· Samantha J. Klasfeld, Ph.D. · Tutorial · 4 min read
Biobank Intro Series: All of Us Observational Data (Part I)
Finding concept IDs for the All of Us Researcher Workbench
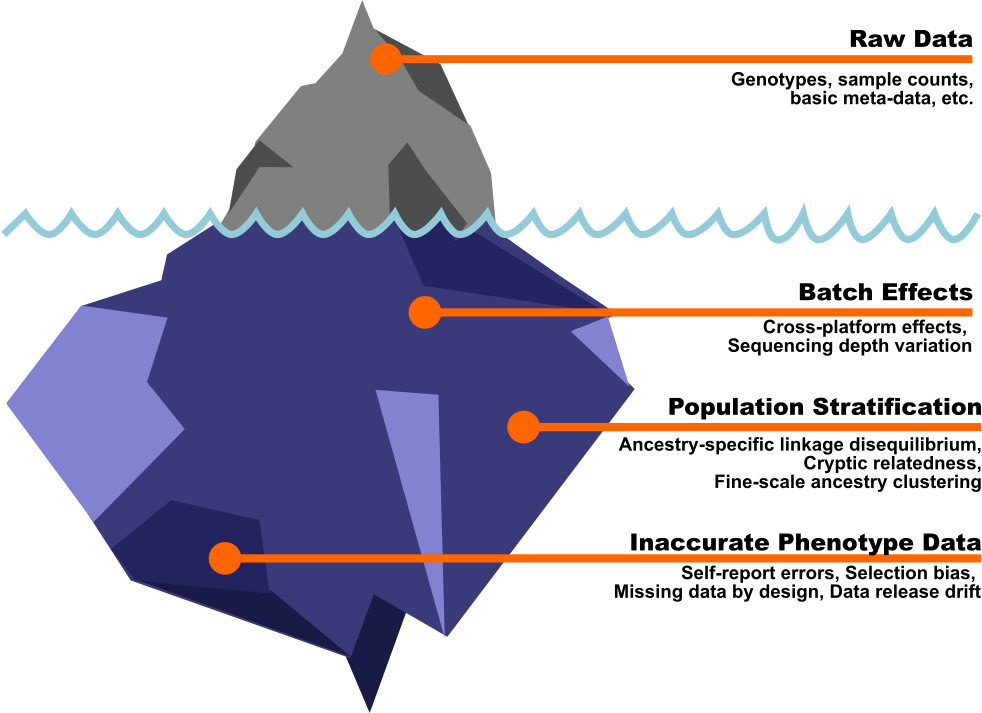
All of Us has returned to the chat (she made her first appearance in the hardware post) and she’s bringing noble intentions and absolutely feral data provenance. UK Biobank is a well-organized remarkable resource, but with >93% European ancestry, it was never designed to represent global genetic diversity. On the other hand, All of Us was explicitly built to oversample underrepresented populations. With almost 50% of participants identifying as a racial or ethnic minority, “All of Us” isn’t just a pun. It’s a mission statement. (I wrote more about why that matters here).
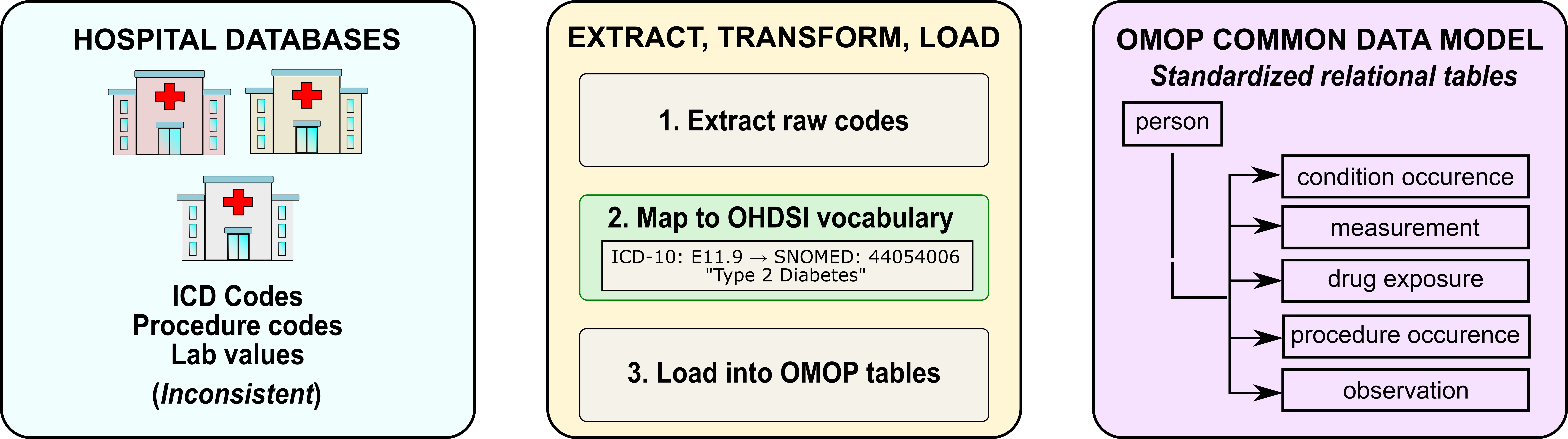
That ambition comes with a data architecture to match. UK Biobank invited ~500,000 people to assessment centers, ran standardized measurements, and stored the results in a single curated dataset with clean field IDs. Even its hospital records flow from a single source: the UK’s publicly funded National Health Service (NHS). All of Us is working with a different reality entirely. It aggregates EHR records from 50+ independent US health systems, enrollment surveys, wearables data, and biosamples, and standardizes all of it after the fact using the OMOP-CDM (Observational Medical Outcomes Partnership Common Data Model) developed by the Observational Health Data Sciences and Informatics (OHDSI) community. What gets recorded in any given participant’s file follows whatever combination of ICD, LOINC, CPT, or SNOMED their health system happened to use.
This is why the workflow feels different. Instead of searching a Showcase and grabbing a field ID, you’re navigating concept IDs, vocabulary mappings, and relational tables: the machinery the OMOP-CDM uses to impose order on that heterogeneity. Concept IDs are the key to unlocking most of what’s available, so let’s start there.
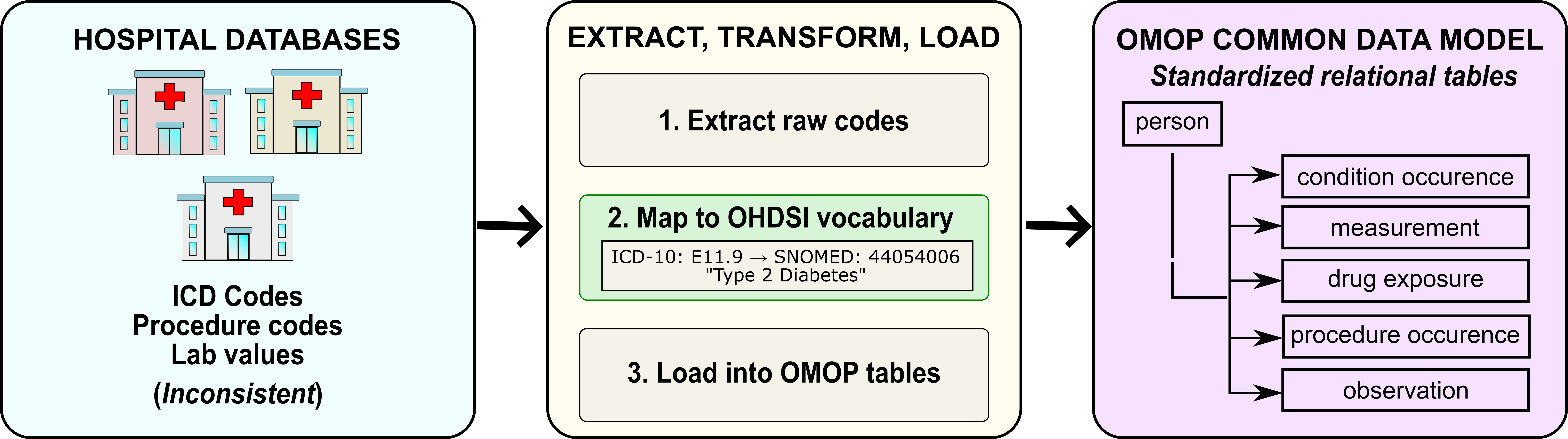
Finding Concept IDs: Three Approaches
Similar to Field IDs, a concept ID is just a number until you know what it maps to. These three tools each give you a different way to look that up, depending on how much context you need.
1. All of Us Data Browser (fastest for quick lookups)
All of Us Data Browser shows what concepts exist and how often they’re used. Search “systolic blood pressure” and you’ll see results under “Conditions” and “Labs & Measurements” - it can be recorded multiple ways.
Click a category to see participant counts and value distributions. For example, 225,000+ participants have concept ID 3004249 (Systolic blood pressure) with source code LOINC 8480-6.
When to use: Quick lookups, checking data availability, getting participant counts.
2. OHDSI Athena Vocabulary (richer metadata)
OHDSI Athena provides concept context such as hierarchies, standard vs. non-standard mappings, and relationships. Searching for “systolic blood pressure” returns about 78,865 items at the time of writing. However, if you look up OMOP-CDM concept ID 3004249 (which you found with the All of Us Data Browser), you’ll see its vocabulary source (LOINC), domain (Measurement), and related concepts.
When to use: Detailed concept information, exploring relationships, understanding vocabulary mappings.
3. Direct SQL queries (most flexible)
Query the CONCEPT table directly to translate medical concepts into IDs:
import pandas as pd
import os
# Use environmental variable for the Controlled Tier
CDR = os.environ['WORKSPACE_CDR']
# Search by name
concept_query = f'''SELECT *
FROM `{CDR}.concept`
WHERE LOWER(concept_name) LIKE '%systolic blood pressure%'
'''
sbp_query = pd.io.gbq.read_gbq(concept_query, dialect='standard')
# Or query a known concept ID directly
concept_id_query = pd.io.gbq.read_gbq(f'SELECT * FROM `{CDR}.concept` WHERE concept_id = 3004249')Use CONCEPT_RELATIONSHIP to explore how concepts relate - for example, which ICD-10 codes map to SNOMED concept 3004249.
When to use: Complex relationship queries, programmatic vocabulary exploration.
What’s Next
Finding concept IDs is the first step, but you need to navigate the relational tables to actually pull observational data. In Part II, we’ll get into the messier, more interesting part.